What we study
Research
We focus on two interconnected research areas: developing scalable computational methods for genomic data, and applying them to biobanks to understand the genetic basis of human disease.
Theme 01
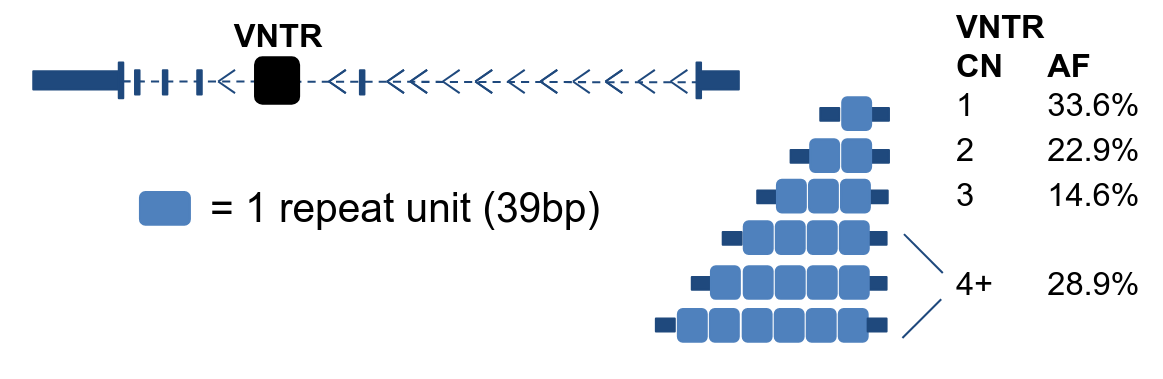
Structural Variation & Disease
Large genomic rearrangements, such as deletions, duplications and inversions are often overlooked in standard GWAS studies, yet account for a substantial fraction of heritable disease risk. We develop methods to detect, phase, and functionally interpret structural variants at biobank scale, integrating them with transcriptomic and proteomic data.
Theme 02
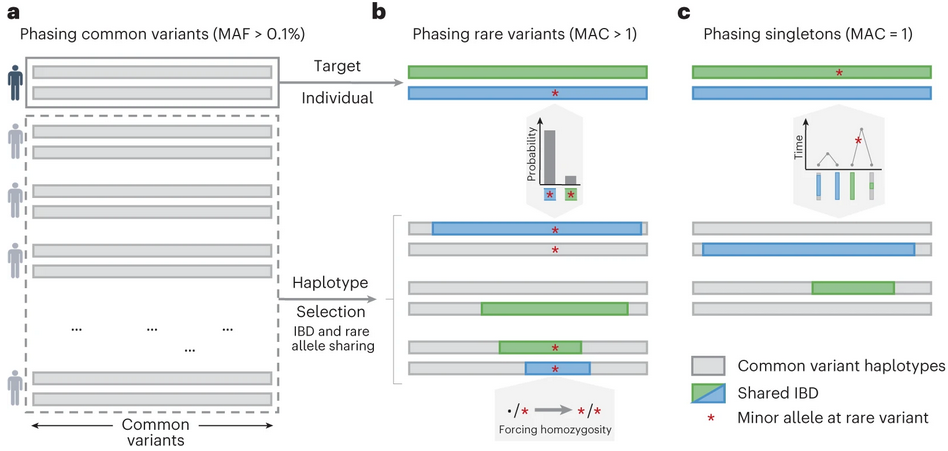
Population Haplotypes & Identity-by-Descent
We analyze genetic relationships within and between populations by identifying shared haplotype segments: regions inherited from a recent common ancestor. This illuminates human population history, enables powerful disease mapping through IBD, and underpins accurate genotype imputation.
Theme 03
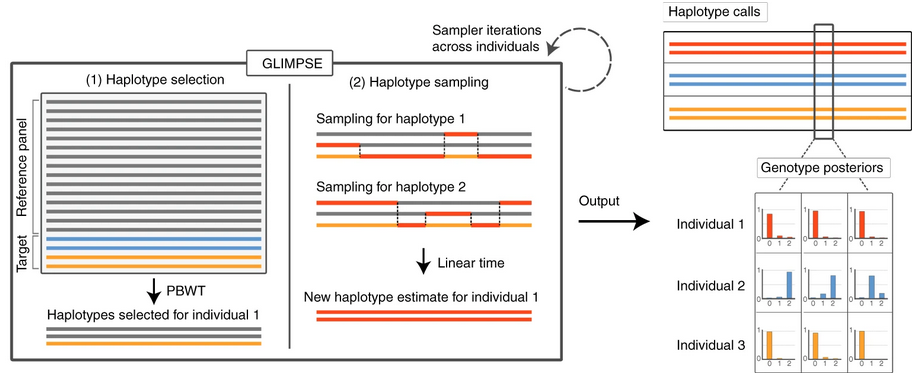
Low-Coverage Sequencing & Imputation
SNP arrays have dominated human genetics for two decades, but low-coverage whole-genome sequencing is rapidly becoming a cost-effective alternative. We develop statistical methods such as GLIMPSE that recover accurate genotypes, including rare variants, from sequencing depths as low as 0.1×.
Theme 04
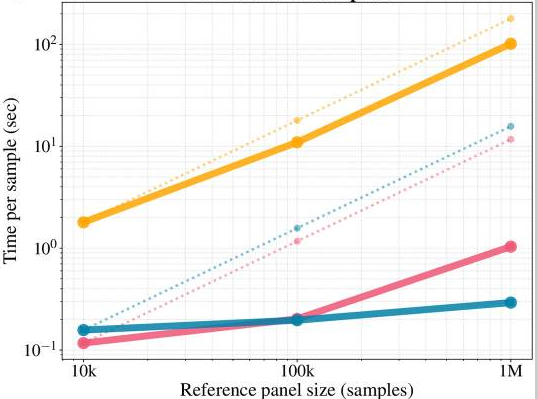
Scalable Computational Algorithms
All our biological questions are ultimately constrained by computation. We design algorithms that scale to millions of genomes. Core methodological themes include Hidden Markov models on haplotype space, Positional Burrows-Wheeler Transform (PBWT), extensions of the Li–Stephens model, and compressed reference panel representations.
