Phasing Singletons: Breaking a 20-Year Barrier in Statistical Genetics
For two decades, population-based phasing methods reported 50% switch error at singleton variants — equivalent to random guessing. SHAPEIT5 changes this, and here's how.
 Simone Rubinacci · 20 July 2023
Simone Rubinacci · 20 July 2023 For two decades, statistical haplotype phasing methods have had a dirty secret: for singleton variants — variants found in only one individual in the entire dataset — they perform no better than random guessing. A 50% switch error rate. Coin-flip accuracy.
SHAPEIT5 changes this. Here’s the story of how, and why it matters.
Why singletons are hard
Standard phasing methods work by identifying shared haplotype segments between individuals (Identity-by-Descent, IBD). If your variant appears in many people, there are many IBD segments to learn from, and phasing is accurate. But if your variant appears in exactly one person — a singleton — there are no IBD segments. There’s nothing to compare against.
The standard approach gives up and phases singletons randomly. This sounds acceptable (singletons are rare, after all), but the problem is that rare variants are increasingly what we care about in disease genetics. Most Mendelian disease mutations are singletons or ultra-rare. And compound heterozygous events — where two pathogenic variants hit both copies of a gene — require accurate phasing to detect.
The SHAPEIT5 approach
SHAPEIT5 uses a coalescent-inspired model that treats each haplotype as a mosaic of segments inherited from other haplotypes in the reference panel. The key innovation: instead of relying only on IBD with other carriers, we use the local haplotype context around the singleton.
Even if only one person carries a rare variant, the haplotype background around that variant is shared with many other people. By modelling the entire haplotype — not just the variant itself — we can infer the most likely phase.
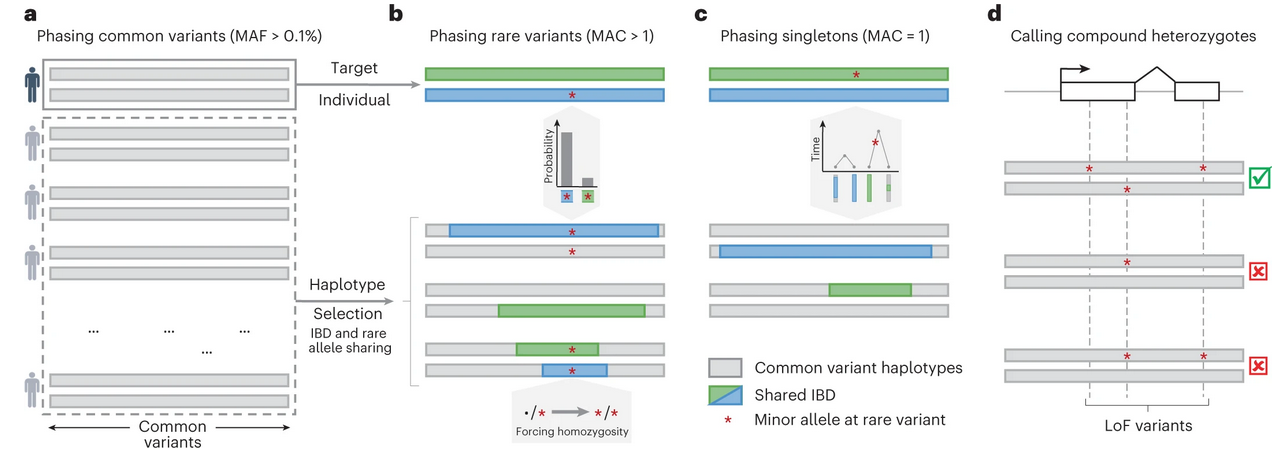
Figure 1: Common and rare variant phasing using SHAPEIT5 and overview of the work.
In practice, SHAPEIT5 achieves:
- < 5% switch error rate for ultra-rare variants (MAF < 0.001%).
- Non-random phasing of singletons — the first method to achieve this without family data.
- < 1% switch error rate for common variants.
Clinical implications
One direct application: detecting compound heterozygous loss-of-function (CH-LOF) events — where one copy of a gene carries a loss-of-function variant on each chromosome. These are a major cause of Mendelian disease.
In the UK Biobank, SHAPEIT5 identified 549 genes with CH-LOF events that would have been missed or misclassified with previous phasing methods. For rare disease diagnosis, accurate phasing is not a statistical nicety — it’s clinically essential.
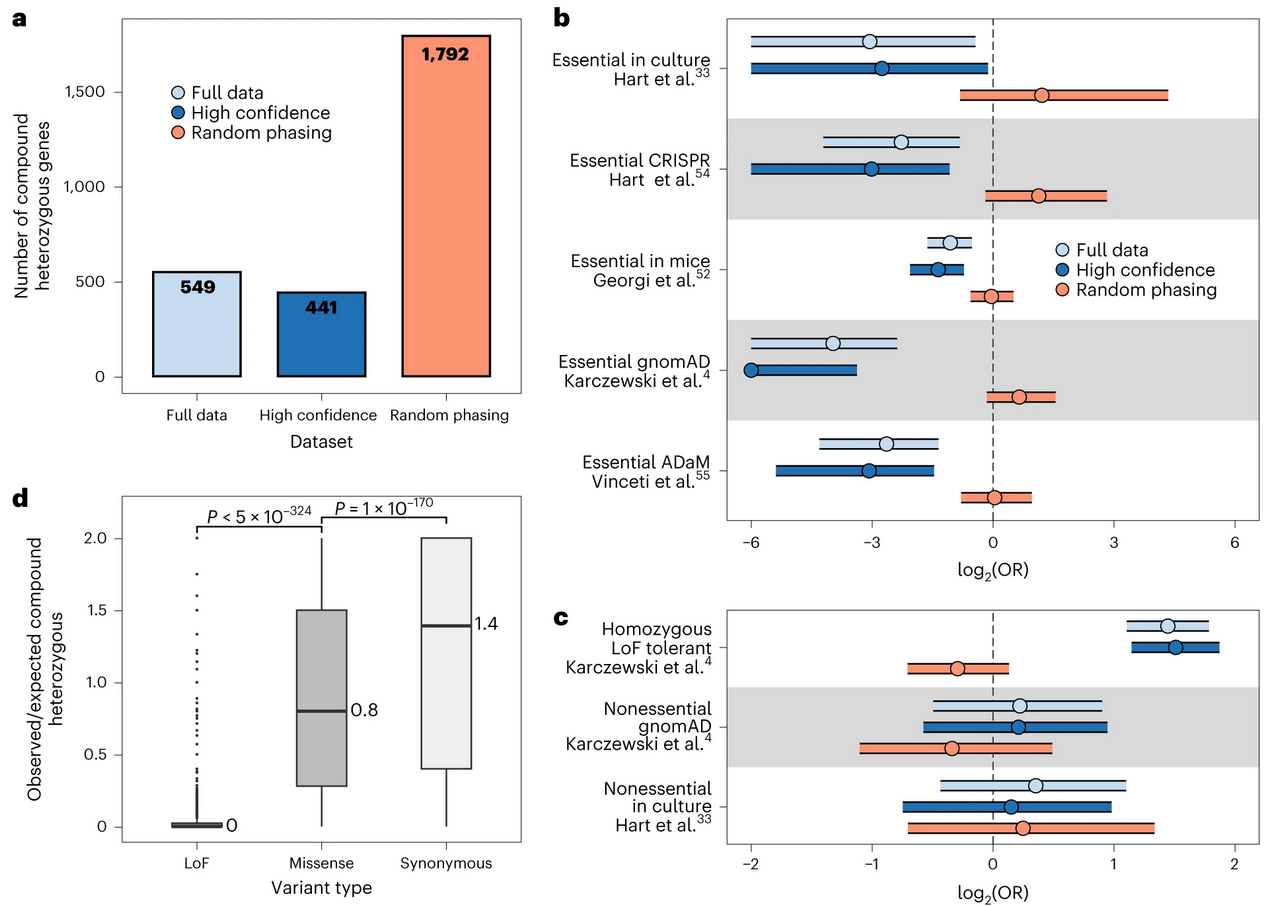
Figure 2: Statistical analysis of our compound heterozygous across different variant annotations
Beyond the UK Biobank
SHAPEIT5 is publicly available and works on any large sequencing dataset. As biobanks grow and whole-genome sequencing becomes standard, the ability to accurately phase rare and singleton variants will only become more important.
Full paper: Hofmeister, Ribeiro, Rubinacci, Delaneau, Nature Genetics 2023. doi:10.1038/s41588-023-01415-w