How We Imputed from 150,000 UK Biobank Genomes for Under $0.1 per sample
The UK Biobank's WGS release was an extraordinary resource — but existing imputation methods couldn't cope at that scale. Here's how GLIMPSE2 was redesigned from the ground up.
 Simone Rubinacci · 15 July 2023
Simone Rubinacci · 15 July 2023 The UK Biobank’s whole-genome sequencing release of 150,119 individuals was one of the most significant events in human genetics in recent years. But it also posed a serious computational challenge: how do you use a reference panel of this size for imputation, when existing methods were designed for panels of tens of thousands?
This post walks through the core ideas behind GLIMPSE2 and why we had to fundamentally redesign the algorithm.
The problem with scaling
Classical imputation methods — including our own GLIMPSE1 — work by first selecting a subset of reference haplotypes for each target individual (using the Positional Burrows-Wheeler Transform), then running a Hidden Markov Model (HMM) over that subset. This approach is fast for moderate-sized panels, but its computational cost scales linearly (or worse) with panel size.
At 280,000 haplotypes, this is simply too slow. A naive approach would have taken years of compute time for the full UK Biobank.
The GLIMPSE2 solution: split-and-merge
The key insight in GLIMPSE2 is that we can split the reference panel into smaller, overlapping chunks, run the HMM on each chunk independently, and then aggregate the posterior probabilities across chunks. This gives us:
- Sublinear scaling: Adding more reference samples increases accuracy, but compute cost grows very slowly.
- Parallelism: Each chunk can be processed independently, enabling efficient use of HPC clusters.
- Accuracy at rare variants: By including locally relevant haplotypes in each chunk, we maintain accuracy even at variants with MAF < 0.1%.
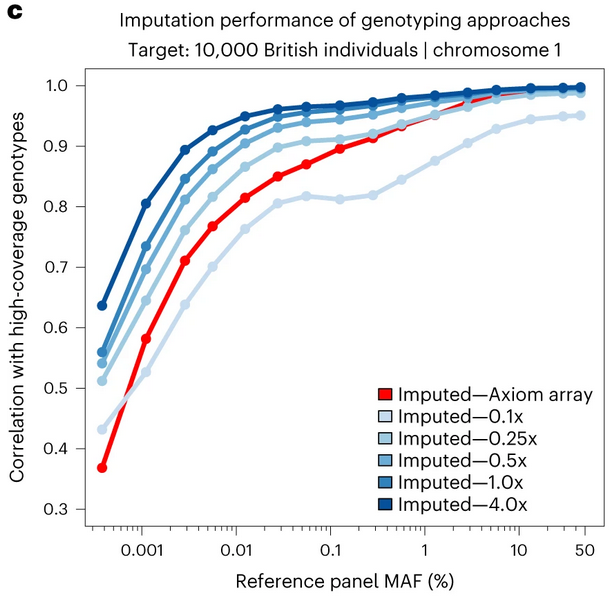
Figure 1: Imputation performance comparison. Note how GLIMPSE2 maintains high correlation with high-coverage genotypes across the frequency spectrum, even at sequencing depths as low as 0.1x.
The result
Using GLIMPSE2, we imputed all 150,119 UK Biobank low-coverage genomes in a matter of weeks, at a compute cost of less than $1 per genome. The resulting dataset — 580 million markers per individual — is among the largest imputed datasets ever produced.
For comparison, the same task with MINIMAC4 (the previous state of the art) would have required roughly 30× more compute. At biobank scale, that difference is the gap between feasible and impossible.

Figure 2: Power in genome-wide association testing across 22 UK Biobank phenotypes. Imputed low-coverage WGS (0.5x and above) consistently outperforms traditional SNP genotyping arrays (Axiom).
What this means for population genetics
The $1-per-genome threshold is significant. It means that low-coverage WGS followed by imputation is now competitive with — and in many ways superior to — SNP genotyping arrays. Arrays typically cost $50–$150 per sample and are limited to pre-defined markers, while lcWGS + imputation provides genome-wide, population-specific variant coverage.
This opens the door to large-scale genetic studies in biobanks and populations where arrays have historically been cost-prohibitive.
Full paper: Rubinacci et al., Nature Genetics 2023. doi:10.1038/s41588-023-01438-3